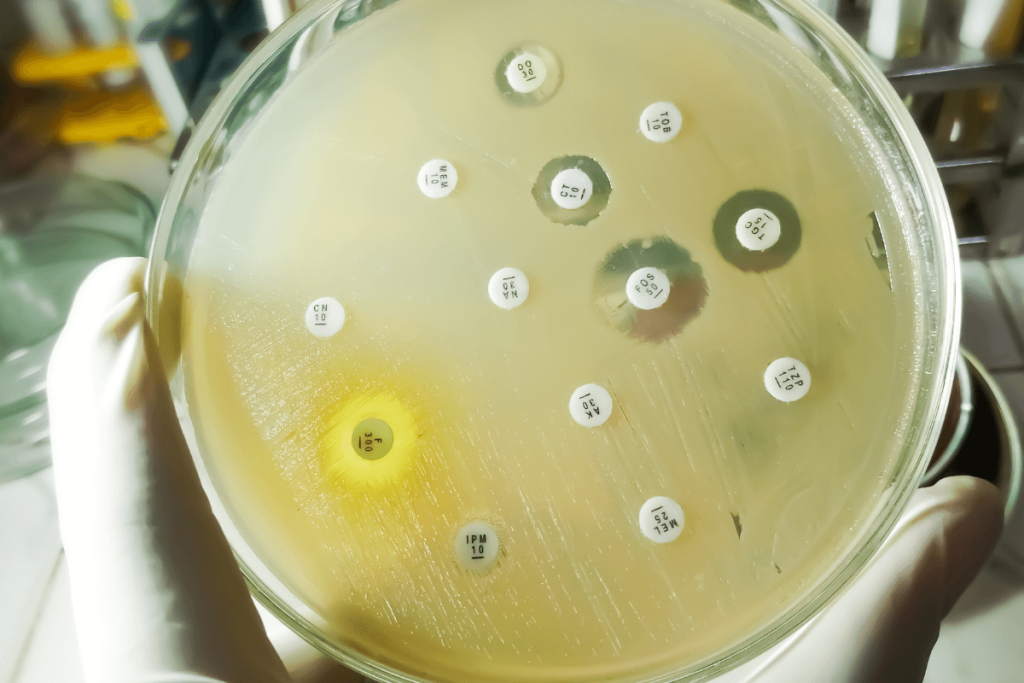
Antimicrobial resistance (AMR) accounts for nearly 9% of global deaths and represents a growing public health threat worldwide. Due to the insufficiency in the development and commercialization of new antibiotics, researchers have turned to the possibility of using antibiotic and non-antibiotic drug combinations as an alternative approach to the management of resistant infections.
Beta blockers, antidepressants, antiepileptics, and nonsteroidal anti-inflammatory drugs (NSAIDs) are examples of many non-antibiotic drugs that have synergistic effects with antibiotics and may improve treatment outcomes against drug-resistant pathogens. Nevertheless, it has also been clinically observed that the application of these drugs can also play a role in the emergence and dissemination of antibiotic resistance in human-related bacteria, probably indirectly via the influence on the gut microbiome.
Experimental studies have shown that various non-antibiotic drugs can enhance the transfer of plasmids carrying antibiotic resistance genes through multiple mechanisms. These involve the rise in production of reactive oxygen species, the SOS response of the bacteria, efflux pumps, and cell membrane permeability. Even some of these drugs can increase bacterial conversion, which raises questions of whether compounds that are intended to treat people will, unwillingly, increase horizontal gene transfer and enhance the resistance to antibiotics in both human and environmental microbiomes.
The aquatic environment is one of the primary factors in the global transmission of antimicrobial resistance. It provides a route to pollutants and resistant bacteria, which may be entered by human populations and animal populations through drinking water, taking a bath, or irrigation in crop fields. Non-antibiotic drugs frequently enter water systems through sewage, pharmaceutical, or industrial effluents and can persist at low yet bioactive concentrations.
Lab experiments suggest that the traces of these drugs can encourage the genetic mutations, propagate the spread of resistance genes, and trigger the plasmid conjugation by oxidative stress. Although this is the case, small-scale studies have been done to assess the impacts of the residual non-antibiotic drug concentrations on antibiotic resistance in natural aquatic systems.
To address this gap, a study that was conducted recently investigated the life cycle of four widely used non-antibiotic drugs, chlorpromazine, diclofenac, diphenhydramine, and fluoxetine in river water, in relation to antibiotic resistance and bacterial community composition. These medications were selected because they belong to various therapeutic classes such as antipsychotics, NSAID substances, antihistamines, and antidepressants. All the compounds were made to be tested at two concentrations: low levels, which were equivalent to the environmental contamination, and high levels, which were equivalent to clinical exposure.
In Lyon, France, the river waters of the Rhone were incubated in eight-day microcosms under control. The use of metagenomic sequencing, 16S rRNA analysis, quantitative PCR, and mass spectrometry was done to examine the bacterial diversity, abundance of genes, and stability of the drug. ANOVA, Kruskal-Wallis, and PERMANOVA were the statistical procedures that provided results on the differences between the control and the treated groups.
The findings indicated that the four non-antibiotic drugs were all chemically stable over the course of the experiment. At very low concentrations, they changed the composition and metabolic activity of bacterial communities, although the extent of each effect varied with compound, dose, and exposure time. Chlorpromazine and diphenhydramine produced the most significant alterations in bacterial structure, whereas fluoxetine enhanced bacterial abundance in general at higher concentrations.
Metagenomic evidence also showed that certain bacterial groups that have resistance genes were enriched selectively. Chlorpromazine and diphenhydramine selectively encouraged the growth of Pseudomonas species that possessed aminoglycoside resistance genes and beta-lactamase genes, whereas fluoxetine encouraged the growth of Sphingobium and Pseudoxanthomonas species that possessed beta-lactamases and efflux pumps. Diclofenac had less pronounced effects, yet it still increased the occurrence of efflux-related resistance genes.
In conclusion, this study demonstrates that even subinhibitory concentrations of non-antibiotic drugs can disrupt microbial ecosystems and promote the selection of bacteria harboring antibiotic resistance genes in aquatic environments. These results indicate that even non-antibiotic popular medications can confer a latent ecosystem contributor to antimicrobial resistance. It is necessary to continuously monitor the environment of such compounds and conduct more studies on the molecular processes better to understand their role in the spread of resistance.
References:Sanchez-Cid C, Vrchovecká S, Dehon E, Wacławek S, Vogel TM. Environmental consequences of anthropogenic pollution: non-antibiotic-drug-driven antibiotic resistance selection in a model aquatic ecosystem. Environ Health. Published online ASAP. doi:10.1021/envhealth.5c00238